|
Use the gentleMACS™Octo Dissociators with Heaters upstream of your single cell genomics analyses.Īutomated, reproducible, hands-off tissue dissociation from fresh, frozen, or OCT-embedded tissue into single cells or singlenuclei.Choose from a selection of automated pre-set protocols and pre-formulated reagents, or create your own protocols and use your reagents of choice. Ready to use programs, in conjunction with Miltenyi tissue dissociation kits that combine mechanical dissociation and enzymatic treatment, yield single-cell suspensions with excellent viability and preserved epitope integrity. Tissue dissociation is performed in a closed and sterile system. Miltenyi Biotec gentleMACS™Octo Dissociators with Heatersįully automate and standardize your tissue dissociation or homogenization of up to eight samples simultaneously. The Chromium Controller also allows for long-range whole genome/exome applications for haplotype phasing & structural variation information, as well as de novo sequencing. Or examine copy number variation of up to 40,000 cells in parallel. Combine single cell transciptomics with single cell chromatin accessibility for a multi-omics approach. Single cell transcriptomics applications include gene expression profiling, simultaneous gene and cell surface protein expression profiling, gene expression CRISPR screening, immune profiling, simultaneous immune & cell surface protein profiling, and immune profiling & antigen specificity. The 10x Genomics Chromium Controller performs high-throughput, parallel partitioning, and automated barcoding for powerful RNA, DNA, and epigenomics sequencing applications for up to 80,000 single cells. Pre-designed and custom panels available. Compatible with fresh or frozen cells or tissue, as well as methanol fixed cells. The Tapestri platform can be used to detect rare subclones (down to 0.1%), identify true mutation co-occurance in clonal populations, resolve zygosity, and reconstruct phylogenetic lineages. Detect SNVs and CNVs in the same cell, or DNA and protein for a multiomics approach. Run up to 8 samples in parallel and capture from 500-10,000 cells per sample.Įxplore mutational diversity on a cell-by-cell basis using the Tapestri single cell targeted DNA-Seq platform.   
Reduce errors from manual pipetting and minimize technical variation to generate consistent and reproducible single cell gene expression results across experiments, users, and sites. The Chromium Connect automates 10x Genomics’ scRNA-Seq assay, going from single cells to sequencing-ready libraries with minimal sample handling using an integrated and validated solution. Up to 7.5 Gb output, 25M reads, and 2x150 bp read length. It features cost-efficient sequencing, even for low numbers of samples. The MiniSeq system delivers the power and confidence of proven Illumina next-generation sequencing technology in an accessible sequencing solution. Up to 15 Gb output, 25 M reads, and 2x300 bp read length. 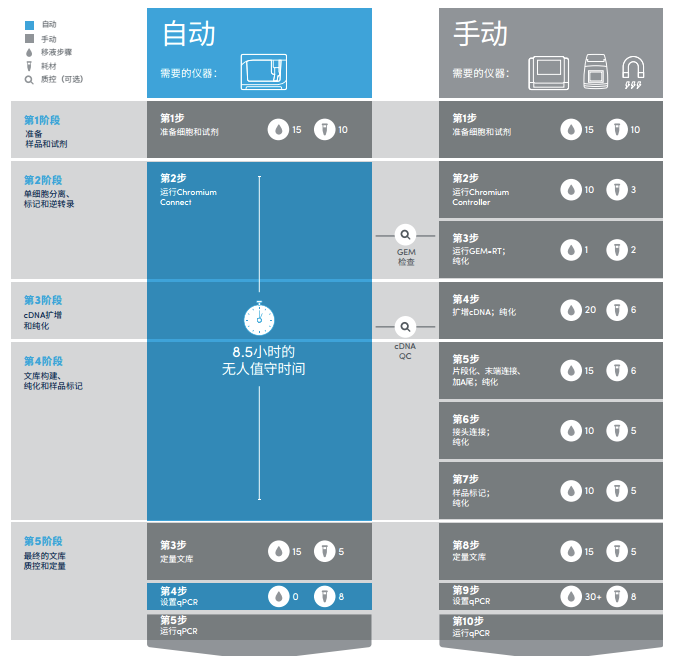
The MiSeq is best suited for whole genome sequencing of smaller model organism genomes, medium to small targeted panels, amplicon sequencing, 16S metagenomics, quality control of libraries, and more. The MiSeq system offers the first end-to-end sequencing solution, integrating cluster generation, amplification, sequencing, and data analysis into a single instrument. Up to 120 Gb output, 400 M reads, and 2x150 bp read length. Two flow cell configurations allow for tunable outputs. The NextSeq 500 system delivers the power and flexibility to carry out applications such as whole genome, exome, transcriptome, and methylation sequencing, among others. Up to 3000 Gb output, 10 B reads, and 2x150 bp read length. Four flow cells are available, as well as the NovaSeq XP workflow that allows for the loading of individual lanes with different library types. Patterned flow cell technology generates an unprecedented level of throughput for a broad range of sequencing applications. The NovaSeq 6000 system offers flexibility for virtually any sequencing method, genome, and scale.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
